Welcome !
By pf2431
Dec. 16, 2020

Discover the PF2431 haplogroup through its history, genealogy and science! Many years of research and discovery have made it possible to know more. These few web pages aim to expose them.

Discover the PF2431 haplogroup through its history, genealogy and science! Many years of research and discovery have made it possible to know more. These few web pages aim to expose them.

A very important publication which inserts in the haplogroup tree all the ancient DNA samples, we find the samples of Ifri n’Amr (IAM4, IAM5 and IAM7):
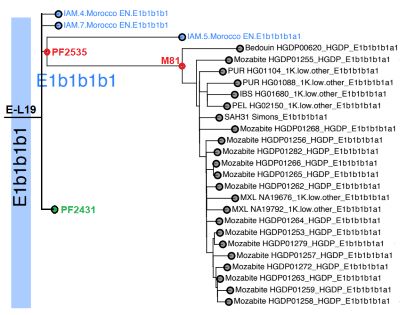
IAM4 and IAM7 are E-L19* (positive for L19, negative for PF2431 and M81)
IAM5 is PF2535* (positive for PF2535 and negative for M81)
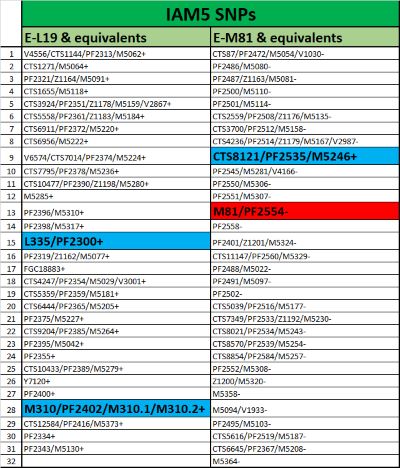
Here M310.1 is equivalent to L19.
Of the 84 snps equivalent to M81, only one was found positive in the old IAM5 dna. It should be remembered that 30,933 snps were found to be positive while each human being has between 3,000 and 5,000 positive snps. Of these 30,933 snps, nearly 28,000 are false positives, it was suspected that PF2535 was the same.
All of the AMIs if related to each other, one would be inclined to think that this small group as a whole had identical results and that they were all L19*. I remind you that all the samples from the Tafoghalt site are all E-M78*, similarly for those from Raqefet Cave (Natufian) which are all E-Z830*. I am quoting what IAM study author R. Fregel said about IAM5 and which goes in this direction:
"IAM.5 only showed one derived SNP, while the remaining were ancestral. We used this evidence to propose that the lineage present in IAM was ancestral to modern E-M81. And it makes sense, IAM samples are around 7,000 years old, while coalescence age for E-M81 is much younger.
Regarding the use of low-coverage data, you have to be careful when analyzing the data. We always classify the mutations taking into account if they can be related to DNA damage (C to T, G to A). The derived SNP in 557 can't be produced by cytosine deamination, so we considered it. Are the mutations that make no sense attributable to DNA damage?"
Which explains why this sample in the article is defined L19*.
In conclusion: What should be remembered is that the majority of the North African population belongs to the L19 (PF2431+M81) sub-haplogroup which was discovered in its ancestral state in skeletons discovered in North Africa and dated at carbon 14 of approximately 7000 years old. It is therefore endogenous to North Africa and has two branches PF2431 and PF2535. It is hoped that other known ancient sites will be studied to gain more information on its history and migrations.

A recent archaeological study conducted in Pompeii has unveiled new perspectives on the daily lives of its inhabitants before the catastrophic eruption of Mount Vesuvius in 79 AD. Excavations have uncovered two skeletons, identified as samples I3682 and I3691, belonging to a woman and a man who had sought refuge in a small room. This exceptional discovery has captured the attention of the global scientific community, as it offers a poignant glimpse into the final moments of these individuals and their struggle for survival.
Preliminary analyses of the samples suggest that both victims took refuge in this room to protect themselves from volcanic ash and debris. The position of the bodies and the objects found nearby depict an emotional scene: the woman, lying on a bed, held a small treasure containing coins and jewels, while the man stood by her side. These material elements highlight the emotional value attached to these objects, even in the most tragic moments of their lives.
Genetic analyses conducted on these samples have revealed fascinating information about the origins and ancestry of the individuals. Sample I3682 carried the haplotype E-Y10561, while sample I3691 carried the haplotype E-L19 (xM81, Y141598, PF2440). These genetic markers are crucial for researchers studying the E-PF2431 haplogroup, providing valuable insights into the ancestral origins of the individuals and their possible familial connections.
The E-PF2431 haplogroup is a major genetic marker used to trace human migrations and understand the ancestral links between different populations. The identification of haplotypes E-Y10561 and E-L19 in the Pompeii individuals sheds new light on the genetic diversity of this ancient population and its connections with other human groups. These discoveries enrich our understanding of genetic evolution and interactions between populations over the ages.
According to research published in Nature Communications and Current Biology, the E haplogroup is one of the oldest and most diverse in Africa and Eurasia. The E-Y10561 and E-L19 subgroups are particularly prevalent in Mediterranean populations, suggesting migrations and genetic exchanges between different regions over millennia.
This study represents a significant advancement in our understanding of the eruption of Mount Vesuvius and its impact on the inhabitants of Pompeii. It also demonstrates the potential of genetic analyses to reveal details about the history and ancestry of ancient individuals. Researchers hope that these findings will inspire future research and contribute to a better understanding of our common genetic heritage.
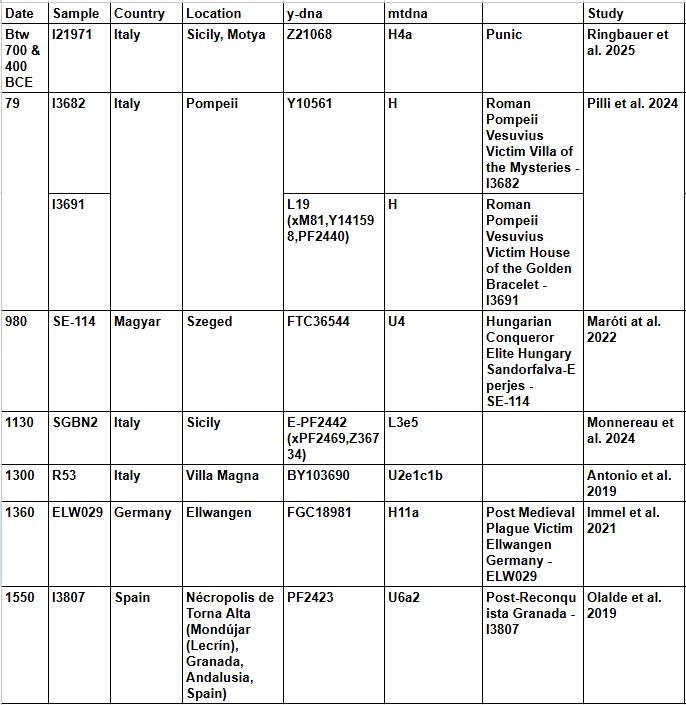
The presence of these haplogroups in the mentioned locations reflects episodes of expansion, migration, and Mediterranean mixing, with a strong North African (Berber) component that has left a lasting genetic legacy across Southern Europe and beyond.
